We use single and paired recordings from cells in flat mounted mouse retina to study visual processing. Cells are recorded either in cell-attached mode to measure spike output or in whole cell voltage-clamp mode to reveal synaptic conductances. During recordings we stimulate the retina with patterns of light projected onto the photoreceptors using a custom digital light projection system. Paired recordings measure signal transfer between synaptically-connected cells. In some experiments, we target specific cell types using a fluorescence signal measured with two-photon illumination. We are able to target cells in this manner while maintaining physiological light responses.
Anatomical circuit mapping
We are developing new strategies to map the circuitry of the mouse retina at the level of individual synaptic connections between molecularly and physiologically identified cell types. We employ techniques such as targeted RNA manipulations, immunohistochemistry and confocal microscopy to quantify synaptic connectivity.
Modeling is a key component of our work as we seek to place our physiological and anatomical measurements in the context of full retinal circuits. We create our own models in the lab, and we collaborate with theorists in other labs. Our modeling efforts fall into two main categories:
1) Models that predict the spike output of retinal ganglion cells (the output cells of the retina) given arbitrary spatiotemporal patterns of light. We use a combination of anatomical and physiological measurements to create “bottom-up” receptive field models that have predictive power.
2) Models that explore the distribution of information across cell types and the dynamics of the representation of visual information. The rich content of the visual world in encoded in ~20 parallel ganglion cell channels. What rules govern how visual information is distributed among these channels? How do these rules depend on the statistics of the visual scene?
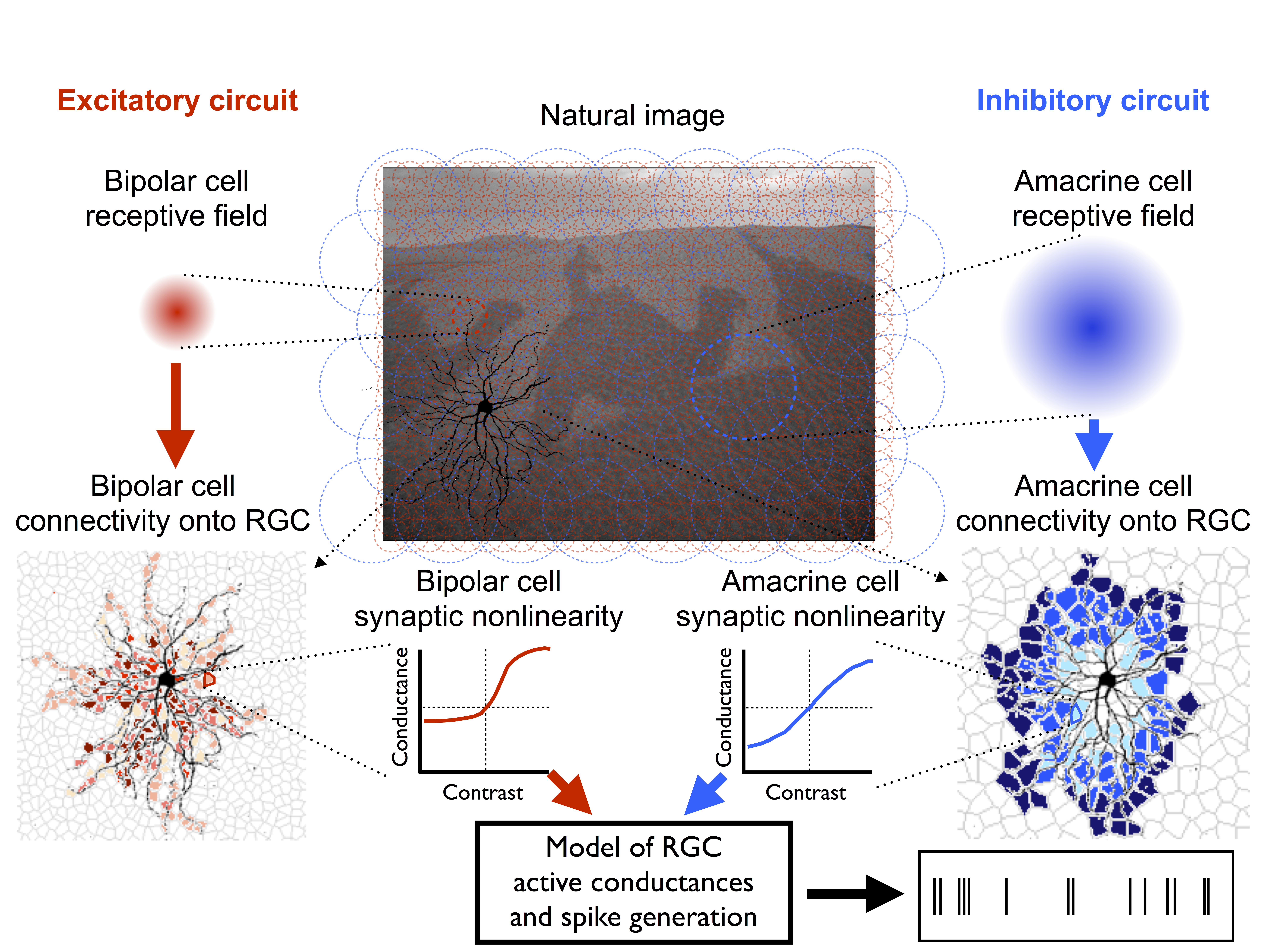
Intraoccular injections
Intracerebral injections
Single cell transcriptomics
Mouse psychophysics